This project couples the unique capabilities of Free Air Concentration Enrichment (FACE) technology, which provides controlled elevation of ozone in open-air at field scale, with the power of the vast genetic resources in maize and RNA-Seq. It will provide a foundation for crop improvement by:
(i) quantifying genetic variation in response to elevated ozone among 200 inbred and 100 hybrid maize lines in the field;
(ii) using high-throughput phenotyping of ozone impacts on maize growth, senescence, leaf metabolism and reproductive processes, to identify traits that correlate with yield loss;
(iii) identifying the genes and gene networks underpinning the ozone response in the most extreme tolerant and susceptible lines, and their hybrids, by integrating RNA-seq expression analysis and detailed physiological analysis in inbred and hybrid maize;
(iv) developing, or identifying existing, biparental populations derived from tolerant and sensitive parents to identify QTLs and eQTLs for ozone tolerance; and
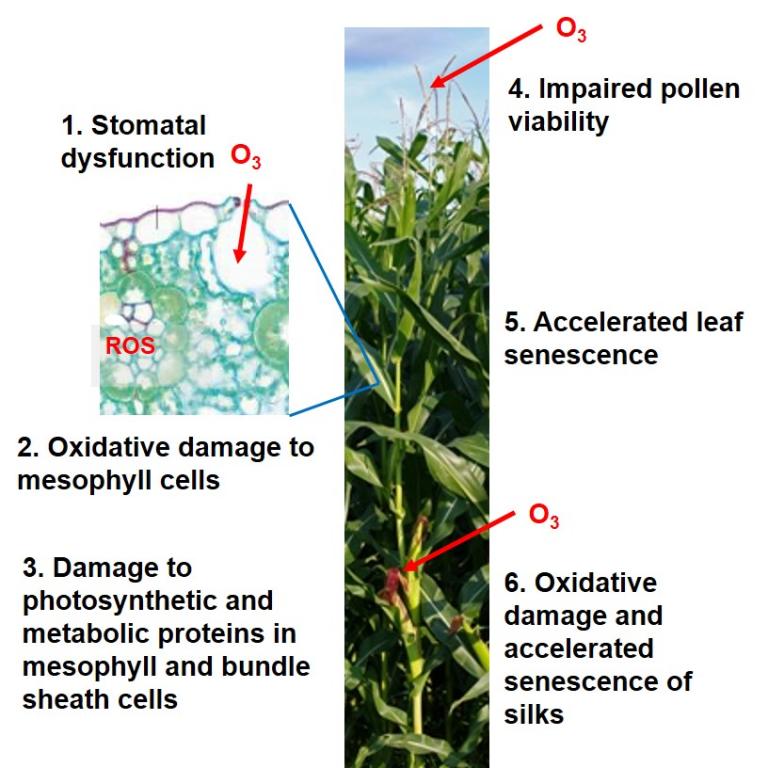
(v) assessing crosstalk between ozone and biotic stress response gene networks in maize. This work will address key mechanistic hypotheses about how oxidative stress leads to transcriptional reprogramming of antioxidant and carbon metabolism, as well as hormone, senescence and defense pathways–integrating the role of tissue specific responses in the stomatal complex, bundle sheath, mesophyll, pollen and silks.
This multifaceted approach is essential because multiple physiological drivers of yield are sensitive to oxidative stress from ozone exposure. Consequently, oxidative stress tolerance is undoubtedly a complex, polygenic trait. But, the broad application of quantitative genetic tools coupled to RNA-seq expression profiling, biochemical and physiological analyses of diverse germplasm makes the challenge of discovering the foundation for ozone tolerance tractable for the first time.